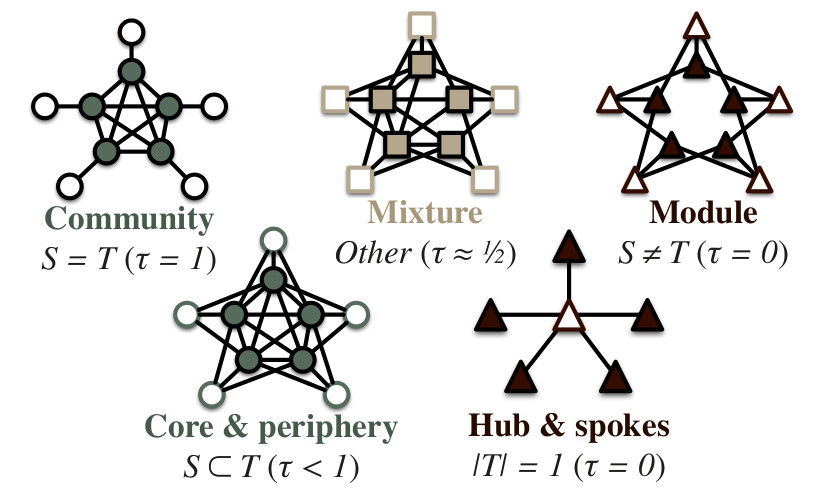
Research tool nets-nodegroups implements the node group extraction framework for network analysis and introduces the group type parameter Tau for researching and exploring node group structures of various networks. The framework is capable of identifying and sequentially extracting significant node group structures, such as:
- communities
- modules
- core/periphery
- hubs & spokes
- and many similar structures
Details of the algorithm and framework are published in:
- L. Šubelj, N. Blagus, and M. Bajec, “Group extraction for real-world networks: The case of communities, modules, and hubs and spokes,” in Proc. of NetSci ’13, 2013, p. 152.
- L. Šubelj, S. Žitnik, N. Blagus, and M. Bajec, “Node mixing and group structure of complex software networks,” Advs. Complex Syst., vol. 17, 2014.
The adopted network node group extraction framework extracts groups from a simple undirected graph sequentially. An optimization method (currently random-restart hill climbing) is used to maximize the group criterion W(S,T) and extract group S with the corresponding linking pattern T. After extraction edges between S and T are removed and the whole process repeated on the largest weakly-connected component until the group criterion W is larger than expected on a Erdös-Rényi random graph.
Open source project:
- home: http://gw.tnode.com/nets-nodegroups/
- github: http://github.com/gw0/nets-nodegroups/
- technology: C++, SNAP library
- citation:
Usage
First compile the tool to get the executable nodegroups (see below). To use it you need your file with graph edges and specify parameters you want to tweak:
-o: prefix for all file names (simplified usage) (default:graph)-i: input file with graph edges (undirected edge per line) (default:graph.edgelist)-l: input file (optional) with node labels (node ID, node label) (default:graph.labels)-og: output file with ST-group assignments (default:graph.groups)-os: output file with only ST-group extraction summary (default:graph.groupssum)-n: number of restarts of the optimization algorithm (default: 2000)-sm: maximal number of steps in each optimization run (default: 100000)-sw: stop optimization if no W improvement in steps (default: 1000)-ss: initial random-sample size of S ant T (0=random) (default: 1)-fn: finish after extracting so many groups (turn off random graphs) (default: 0)-fw: finish if W smaller than top percentile on random graphs (default: 1)-rg: random graphs (Erdos-Renyi) to construct for estimating W (default: 500)-rn: random graph restarts of the optimization algorithm (default: 10)-rf: random graph re-estimation of W if relative difference smaller (default: inf)
Example to read from graph.edgelist and output to graph.groups and graph.groupsreport:
$ ./nodegroups -o:graphSame example but extract only first 12 groups (ignoring estimated W on random graphs):
$ ./nodegroups -o:graph -fn:12Input file graph.edgelist contains undirected graph edges (-i:):
0 1
0 2
1 2
2 3
3 4
3 5
...Output file graph.groups contains extracted node groups S and linking patterns T (-og:):
# NId GroupS GroupT NLabel
0 0 0 foo
1 0 0 bar
2 0 0 foobar
3 1 -1 -
2 -1 1 -
...Output file graph.groupsreport contains a summary of extracted node groups (-or:):
# Graphs: 12 Nodes: 115 Edges: 613
N M N_S M_S N_T M_T N_ST M_ST L_ST L_STc W Tau Mod_S Mod_T Type
115 613 9 36 9 36 9 36 72 25 823.0000 1.0000 0.1352 0.1352 COM
115 582 9 36 9 36 9 36 72 30 818.0000 1.0000 0.1164 0.1164 COM
115 550 10 40 10 40 10 40 80 30 810.0000 1.0000 0.1191 0.1191 COM
...Description of columns:
N: number of nodes left in graphM: number of edges left in graphN_S: number of nodes in subgraph on group SM_S: number of edges in subgraph on group SN_T: number of nodes in subgraph on linking pattern TM_T: number of edges in subgraph on linking pattern TN_ST: number of nodes in subgraph on intersection of S and TM_ST: number of edges in subgraph on intersection of S and TL_ST: number of edges L(S,T) between groups S and TL_STc: number of edges L(S,Tc) between groups S and complement of TW: group critetion W(S,T)Tau: group type parameter Tau(S,T)Mod_S: modularity measure on group SMod_T: modularity measure on linking pattern TType: human name for group type parameter Tau:COM: community (S = T)MOD: module (S intersection with T = 0)HSD: hub&spokes module (module and |T| = 1)MIX: mixture (otherwise)CPX: core/periphery mixture (S subset of T or T subset of S)
Exit status codes:
-1: error, file not found or crash0: success, groups extracted1: no groups extracted
Build
You will need to compile the source code to get a working tool. This should work on any platform using any standard C++ compiler (eg. GCC, Visual Studio), but it has only been extensively tested in the following environment.
First you need to download source code of net-nodegroups:
$ git clone http://github.com/gw0/nets-nodegroups.git
$ cd ./nets-nodegroupsDownload and compile SNAP library from console using GCC:
$ apt-get install build-essential g++
$ wget http://snap.stanford.edu/releases/Snap-2.1.zip
$ unzip Snap-2.1.zip
$ mv Snap-2.1 snap
$ cd ./snap
$ make all
$ cd ..Compile net-nodegroups from console using GCC:
$ cd ./src
$ make allExecutable file is located in ./src/nodegroups.
Feedback
If you encounter any bugs or have feature requests, please file them in the issue tracker, or even develop it yourself and submit a pull request on GitHub.
License
Copyright © 2014-15 gw0 [http://gw.tnode.com/] <>
This code is licensed under the GNU Affero General Public License 3.0+ (AGPL-3.0+). Note that it is mandatory to make all modifications and complete source code publicly available to any user.